The rapid evolution of computer-aided detection (CAD) and computer-aided diagnosis have resulted in significant improvements in the field of medical imaging. Medical imaging is a technique used to capture images of the interior of the body (organs and tissues) for clinical analysis, medical diagnosis, and intervention. These improvements are the result of the need for more precise treatments, increasingly sophisticated computer systems, and the availability of superior compute power.
A critical component for medical imaging is segmentation accuracy. A marginal segmentation error would lead to unreliable results, thus potentially being rejected for clinical settings. Segmentation is particularly helpful in the diagnosis and treatment of brain tumors. Splitting the tumor into smaller sections allows doctors to better understand the tumor's size and shape. This in turn helps medical professionals develop effective treatment plans. Segmenting brain tumors could help reduce margin of error. Such errors could have catastrophic effects on patients' lives.
Image Segmentation: Challenges
Presentation of tumors typically exists within heterogeneous tissues. Heterogeneous tissues look very different from one area to the next, making it hard to determine if the tissues being examined are normal or abnormal. Furthermore, tumors have irregular shapes and sizes that change over time.
Effective and reliable Brain tumor segmentation (BTS) is difficult to achieve due to reasons like:
- Developers working in medical imaging face technical issues, such as large file sizes, image accuracy, image heterogeneity across the organization, and testing and deploying medical image applications, which are time-consuming processes.
- Lack of information about a tumor's specific characteristics makes it difficult to train an AI's deep neural networks to differentiate between healthy and tumorous cells.
- Developers looking for options to deploy medical imaging AI workloads are technologically constrained by:
- The high cost of GPU memory footprint, lack of sample code/solutions to use as a base for their applications, so as not to have to start from scratch.
- Developers are bound by the performance that popular frameworks like TensorFlow and PyTorch offer.
- Performance limitations of popular frameworks like Keras, TensorFlow or PyTorch
- The lack of an easy to use, accessible and affordable platform to validate their solutions and identify the best hardware all makes Clinical and Healthcare tech decision-makers who are direct consumers hesitant to invest.
U-Net Architecture: A Model for optimized Image Segmentation
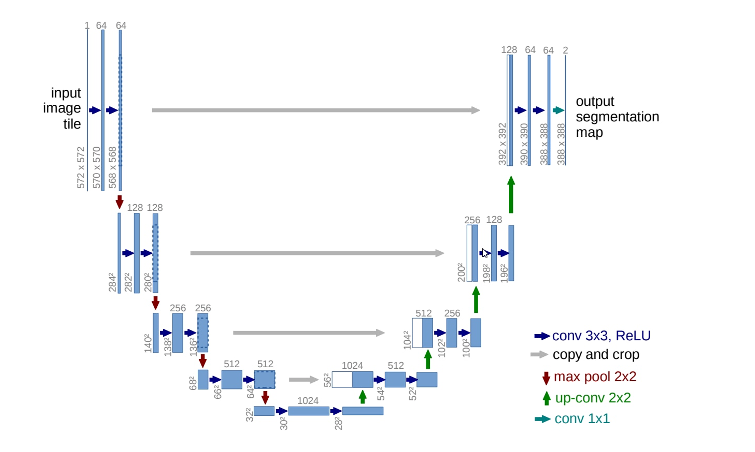
U-Net architecture is a fully convolutional network first implemented in Biomedical image segmentation, which requires fewer training images to yield more precise segmentations. This consumes less time and less compute power. U-Net architecture works by adding the usual contracting network of successive layers, where pooling operations are replaced by upsampling operators. These layers increase the resolution of the output. A successive convolutional layer learns to assemble a precise output based on this information. Up until a certain point, U-Net output is identical to the original contracting network.
The U-Net Sorensen DICE Coefficient uses a new method called discriminative cell classification features (DCCF). It first builds an optimal classification model from DCCF with support vector machine (SVM) and then trains the DNN for tumor segmentation with supervised learning. A major advantage of this approach is that it maintains accuracy and higher-quality precision of tumor segmentation than existing methods. Which use only classifiers or deep neural networks. It also provides more reliable results for clinical imaging and diagnosis using complementary techniques.
Numerous studies have demonstrated that U-Net architecture excels in biomedical imaging segmentation, producing very good results with as little as 50 training images and very reasonable training time.
Fig-1: U-Net Architecture Diagram
Source: U-Net architecture design | Download Scientific Diagram (researchgate.net)
Intel OpenVino optimized Keras- TensorFlow Model for Brain Tumor Segmentation
This is where Intel OpenVino U-Net Sorensen DICE Coefficient Technologies come into play. They offer methods to boost accuracy and reduce false positives without compromising computational efficiency. They do this via sophisticated XPU accelerated deep learning algorithms.
The Intel team optimized the Keras/TensorFlow model with OpenVINO and compared the performance between these models. Where the OpenVINO models showed a significant inferencing performance improvement. OpenVINO optimizations also make the models compatible and inference runs optimally on any Intel architecture. Because the OpenVINO Inference Engine uses built-in plug-ins, eliminating the need for any additional programming. Interoperability also allows for heterogeneous processing, such as running different U-Net topology layers on different hardware using the HETERO command to reduce latency, or running the model on multiple devices using the MULTI command to increase throughput. Compatibility gives developers the ability to choose the right hardware, but in practice it would be difficult for any developer to have a large lab with such a set of testing equipment.
BTS was provided on Intel DevCloud for Edge as an example application that provides sample code for model optimization. Intel’s DevCloud for Edge provides a large cluster of hardware for free. Enables Developers to run the BTS model on different hardware (edge nodes) and compare performance with Keras/Tensorflow and segmentation results with ground truth to calculate Sorensen-Dyes coefficient. This application can act as a reference application or base code that allows developers to implement OpenVINO and DevCloud architecture features in their workload. Intel's DevCloud for the edge gives developers the flexibility to become interoperable and choose the right hardware from the range of options.
Conclusion
Brain Tumor Segmentation is one of the critical use cases for validating Image Segmentation accuracy. Brain tumor segmentation is not an easy task to undertake. But with the right tools and technology, it can be made much easier. Multiple iterative studies have revealed the improved inferencing performance through the Intel OpenVino U-Net Sorensen DICE Coefficient Technologies. Coming into play - Intel's tools offer methods to boost accuracy and reduce false positives without compromising computational efficiency. Intel tools do this with sophisticated XPU accelerated deep learning algorithms.
Intel developed a tool called OpenVino U-Net Sorensen DICE Coefficient Technologies based on the OpenVino Model server. For more information about U-Net and Intel’s whole suite of tools, refer to the Brain Tumor Segmentation Reference Implementation for working models. Visit the Intel Devcloud for Edge and AI Software Development tools documentation.
You must be a registered user to add a comment. If you've already registered, sign in. Otherwise, register and sign in.