- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi,
I have my deep learning model running well on CPU and GPU via OpenVINO. When I specify the Neural Compute Stick 2 as my inference engine, however, I get incorrect prediction from NCS 2. I am using Anaconda command prompt on Windows 10. Could you please help me to sort out this issue? Thank you.
To run the code please unzip the attached file (all the necessary files e.g. input file and IR file are attached).
For inferencing on CPU:
python CNN.py -i X_one -m frozen_graph.xml -d cpu
For inferencing on NCS 2:
python CNN.py -i X_one -m frozen_graph.xml -d MYRIAD
Link Copied
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
Can you share with us the output you are getting with MYRIAD and CPU.
Best Regards,
Surya
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Surya,
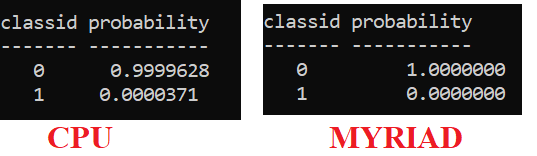
Thank you. The output that I am getting:
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
Can you try with some other input to ensure that the category with higher probability is same for both CPU and MYRIAD.
Best Regards,
Surya
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Surya,
I tried with some other inputs, the estimated probability by CPU were correct for all input. However, estimated probability by MYRIAD were incorrect. "classid 1" always has the highest probability when I use MYRIAD inferencing.
For instance, for a same input:
CPU:
classid probability
------- -----------
0 0.8982905
1 0.1017096
MYRIAD:
classid probability
------- -----------
1 0.9990234
0 0.0014257
Could you let me know what is problem?
Thanks,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
I am using OpenVino 2020.1.33 and Tensorflow 1.15.2. my code for inferencing:
from __future__ import print_function
import sys
import os
from argparse import ArgumentParser, SUPPRESS
import cv2
import numpy as np
import logging as log
from time import time
from openvino.inference_engine import IENetwork, IECore
import threading
from numpy import load
class InferReqWrap:
def __init__(self, request, id, num_iter):
self.id = id
self.request = request
self.num_iter = num_iter
self.cur_iter = 0
self.cv = threading.Condition()
self.request.set_completion_callback(self.callback, self.id)
def callback(self, statusCode, userdata):
if (userdata != self.id):
log.error("Request ID {} does not correspond to user data {}".format(self.id, userdata))
elif statusCode != 0:
log.error("Request {} failed with status code {}".format(self.id, statusCode))
self.cur_iter += 1
log.info("Completed {} Async request execution".format(self.cur_iter))
if self.cur_iter < self.num_iter:
# here a user can read output containing inference results and put new input
# to repeat async request again
self.request.async_infer(self.input)
else:
# continue sample execution after last Asynchronous inference request execution
self.cv.acquire()
self.cv.notify()
self.cv.release()
def execute(self, mode, input_data):
if (mode == "async"):
log.info("Start inference ({} Asynchronous executions)".format(self.num_iter))
self.input = input_data
# Start async request for the first time. Wait all repetitions of the async request
self.request.async_infer(input_data)
self.cv.acquire()
self.cv.wait()
self.cv.release()
elif (mode == "sync"):
log.info("Start inference ({} Synchronous executions)".format(self.num_iter))
for self.cur_iter in range(self.num_iter):
# here we start inference synchronously and wait for
# last inference request execution
self.request.infer(input_data)
log.info("Completed {} Sync request execution".format(self.cur_iter + 1))
else:
log.error("wrong inference mode is chosen. Please use \"sync\" or \"async\" mode")
sys.exit(1)
def build_argparser():
parser = ArgumentParser(add_help=False)
args = parser.add_argument_group('Options')
args.add_argument('-h', '--help', action='help', default=SUPPRESS, help='Show this help message and exit.')
args.add_argument("-m", "--model", help="Required. Path to an .xml file with a trained model.",
required=True, type=str)
args.add_argument("-i", "--input", help="Required. Path to a folder with images or path to an image files",
required=True, type=str, nargs="+")
args.add_argument("-l", "--cpu_extension",
help="Optional. Required for CPU custom layers. Absolute path to a shared library with the"
" kernels implementations.", type=str, default=None)
args.add_argument("-d", "--device",
help="Optional. Specify the target device to infer on; CPU, GPU, FPGA, HDDL or MYRIAD is "
"acceptable. The sample will look for a suitable plugin for device specified. Default value is CPU",
default="CPU", type=str)
args.add_argument("--labels", help="Optional. Labels mapping file", default=None, type=str)
args.add_argument("-nt", "--number_top", help="Optional. Number of top results", default=10, type=int)
return parser
def main():
log.basicConfig(format="[ %(levelname)s ] %(message)s", level=log.INFO, stream=sys.stdout)
args = build_argparser().parse_args()
model_xml = args.model
model_bin = os.path.splitext(model_xml)[0] + ".bin"
# Plugin initialization for specified device and load extensions library if specified
log.info("Creating Inference Engine")
ie = IECore()
if args.cpu_extension and 'CPU' in args.device:
ie.add_extension(args.cpu_extension, "CPU")
# Read IR
log.info("Loading network files:\n\t{}\n\t{}".format(model_xml, model_bin))
net = IENetwork(model=model_xml, weights=model_bin)
if "CPU" in args.device:
supported_layers = ie.query_network(net, "CPU")
not_supported_layers = [l for l in net.layers.keys() if l not in supported_layers]
if len(not_supported_layers) != 0:
log.error("Following layers are not supported by the plugin for specified device {}:\n {}".
format(args.device, ', '.join(not_supported_layers)))
log.error("Please try to specify cpu extensions library path in sample's command line parameters using -l "
"or --cpu_extension command line argument")
sys.exit(1)
#assert len(net.inputs.keys()) == 1, "Sample supports only single input topologies"
#assert len(net.outputs) == 1, "Sample supports only single output topologies"
log.info("Preparing input blobs")
input_blob = next(iter(net.inputs))
out_blob = next(iter(net.outputs))
net.batch_size = len(args.input)
# Read and pre-process input images
#print(type(input_blob))
#print(net.inputs.values())
print(str(args.input[0]))
#X_all = load(str(args.input[0]))
#print(X_all)
n, c, h, w = net.inputs[input_blob].shape
#image = load('C:/Program Files (x86)/IntelSWTools/openvino_2020.1.033/deployment_tools/inference_engine/samples/python/classification_sample_async/1/'+'X_one'+'.npy')
image = load((str(args.input[0])+'.npy'))
image=(image.transpose((0, 3, 1,2)))
images = np.ndarray(shape=(image.shape))
print(n)
print(c)
print(h)
print(w)
for i in range(n):
# image = load('C:/Program Files (x86)/IntelSWTools/openvino_2020.1.033/deployment_tools/inference_engine/samples/python/classification_sample_async/1/'+'X_one'+'.npy')
# print(image.shape)
#image = load('C:/Program Files (x86)/IntelSWTools/openvino_2020.1.033/deployment_tools/inference_engine/samples/python/classification_sample_async/1/'+str(args.input[0])+'.npy')
# image = cv2.imread(args.input)
# if image.shape[:-1] != (h, w):
# log.warning("Image {} is resized from {} to {}".format(args.input, image.shape[:-1], (h, w)))
# image = cv2.resize(image, (w, h))
# image = image.transpose((2, 0, 1)) # Change data layout from HWC to CHW
images = image
# images = image
images.astype('float32')
# print(images.shape)
# print(images[0])
log.info("Batch size is {}".format(n))
# Loading model to the plugin
log.info("Loading model to the plugin")
exec_net = ie.load_network(network=net, device_name=args.device)
# create one inference request for asynchronous execution
request_id = 0
infer_request = exec_net.requests[request_id];
num_iter = 10
request_wrap = InferReqWrap(infer_request, request_id, num_iter)
# Start inference request execution. Wait for last execution being completed
request_wrap.execute("async", {input_blob: images[0]})
# Processing output blob
log.info("Processing output blob")
res = infer_request.outputs[out_blob]
log.info("Top {} results: ".format(args.number_top))
if args.labels:
with open(args.labels, 'r') as f:
labels_map = [x.split(sep=' ', maxsplit=1)[-1].strip() for x in f]
else:
labels_map = None
classid_str = "classid"
probability_str = "probability"
for i, probs in enumerate(res):
probs = np.squeeze(probs)
top_ind = np.argsort(probs)[-args.number_top:][::-1]
print("Image {}\n".format(args.input))
print(classid_str, probability_str)
print("{} {}".format('-' * len(classid_str), '-' * len(probability_str)))
for id in top_ind:
det_label = labels_map[id] if labels_map else "{}".format(id)
label_length = len(det_label)
space_num_before = (7 - label_length) // 2
space_num_after = 7 - (space_num_before + label_length) + 2
space_num_before_prob = (11 - len(str(probs[id]))) // 2
print("{}{}{}{}{:.7f}".format(' ' * space_num_before, det_label,
' ' * space_num_after, ' ' * space_num_before_prob,
probs[id]))
print("\n")
log.info("This sample is an API example, for any performance measurements please use the dedicated benchmark_app tool\n")
if __name__ == '__main__':
sys.exit(main() or 0)
When I use MYRIAD for inferencing "classid 1" always have the higher probability. Could you help me sort out the issue?
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
In addition to what Chauhan, Surya Pratap Singh (Intel) suggested, please share how you converted the model and also which data type are you using (FP16 or FP32)?
Regards,
Javier A.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Javier,
Thank you for your response. Tensorflow 1.15.2 was used to build the CNN model for image recognition. The model is then converted by OpenVino Model Optimizer. The data type was specified to be FP16. The model optimizer command that I used:
python mo.py --input_model inference_graph_onelayer_93recall.pb --input_shape [1,400,90,1] --data_type FP16
The model was successfully converted into the IR files. Please let me know if you need any additional information. Thank you.
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
Could you provide us a variety of sample inputs and their respective expected probability result?
Also, make sure all the layers used in your custom model are supported by the MYRIAD plugin.
Additionally, the Myriad Plugin supports the networks specified in this documentation.
Regards,
Javier A.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Thank you Javier.
My model is a simple Convolutional Neural Network (CNN) to classify image. I used the Keras Sequential API, to create and trained the model. I check the link that you send an it seems that all the layers in my model are supported by the MYRIAD plugin.
The model has only three layers and the input is a 1-channel image. This is the code that I used for creating and training the model:
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Convolution2D, MaxPooling2D, Dense, Dropout, Activation, Flatten
from sklearn.model_selection import KFold
Model = Sequential()
Model.add(Convolution2D(filters = 10, kernel_size = 20, activation='relu', input_shape=(400,90,1)))
Model.add(Flatten())
Model.add(Dense(2, activation='softmax'))
Model.compile(loss='binary_crossentropy', optimizer='adam',metrics=[tf.keras.metrics.Recall()])
history=Model.fit(X_trains_main,Y_trains_main,epochs=1000,batch_size=128 ,validation_split=0.1, verbose=1)
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
Can you provide multiple sample inputs (and expected outputs) for both classes (classid 1 and 0)? Like the actual files, just like you did in the original post (like X_one).
Regards,
Javier A.
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Javier,
Please let me know if you need more information or files. Thank you.
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Javier,
Have you been able to look at the new inputs?
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hello,
Could anyone kindly let me know how I can solve this problem?
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
Thank you for your patience. We are still looking into this issue and will get back to you as soon as we have more information.
Best Regards,
Sahira
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Thank you Sahira. I am looking forward to hearing back from you.
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Reza,
This looks like a bug and I have escalated to the Development Team to take a further look.
I will let you know if they find anything!
Best Regards,
Sahira
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Thank you for your time and consideration Sahira. Our team is looking into different edge devices that are in the market now to see which one would be the best for real-time detection. NCS2 shows a fast response. However, the predictions doesn't seem correct. It would be great if you let me know if you could fix the bug.
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
Hi Sahira,
I was wondering if there is any update regarding my problem. Thank you.
Regards,
Reza
- Mark as New
- Bookmark
- Subscribe
- Mute
- Subscribe to RSS Feed
- Permalink
- Report Inappropriate Content
I was wondering if there is any update from the Development Team regarding my problem.
- Subscribe to RSS Feed
- Mark Topic as New
- Mark Topic as Read
- Float this Topic for Current User
- Bookmark
- Subscribe
- Printer Friendly Page